|
12/17/2023 0 Comments Palindromic sequences
Palindromic sequences in the genome are essentially “unstable,” rendering them vulnerable to insertions and deletions. Some examples are TATA box, a core promoter sequence and binding site for transcription-initiating factors 12 TGACGTCA, the cAMP-responsive element that binds B-ZIP proteins and binding sites of cancer-associated TFs such as SMAD3/4 13, 14.įull size image Palindromic variations in disease-associated loci Palindromes and near-palindromes also serve as binding sites for restriction enzymes and transcription factors (TFs). Because of this tendency, they are often responsible for chromosomal translocations, recombinations, and deletions and are associated with the inheritance of many genetic diseases. Palindromic AT-rich repeats (PATRRs) can cause DNA to denature more readily due to weak A–T bonds, thereby increasing the propensity of the sequence to fold into a secondary structure 11. They can alter the DNA replication process and inhibit gene expression by inhibiting ribosomal translocation along the mRNA transcript 9, 10. The sites of cruciform formation are hot spots for DNA breakage and chromosomal translocation 8. 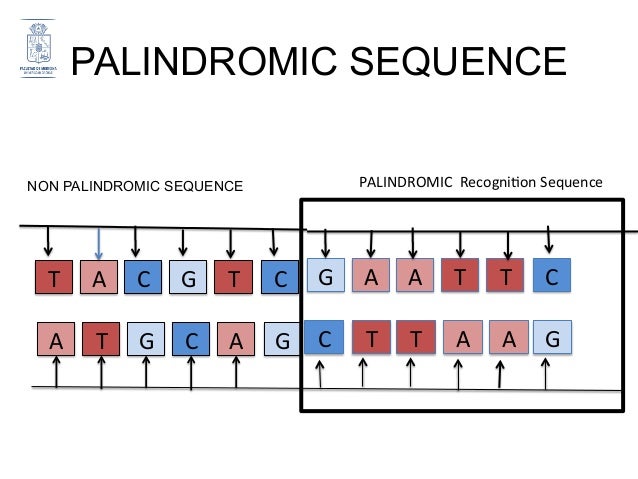
A long palindrome has the unique ability to fold back onto itself to form a secondary structure called a cruciform or hairpin (Fig. Short palindromes, under 50 bp, prevent DNA degradation, while palindromes longer than 50 bp result in mutations and DNA instability 6. They stimulate deletions during DNA replication and interchromosomal recombination between homologous sequences, leading to the loss of intervening sequences in fact, palindromes and near-palindromes account for 83% of deletions and small insertions 4, 5. Palindromes affect various cellular processes, including gene expression, regulation, and gene replication 4. Measuring the abundance, frequency, and location of palindromes across the human genome is critical to understanding their functional roles. Palindromic DNA sequences are prevalent in the genomes of a wide variety of organisms, including humans the functions and implications of such sequences are only beginning to be understood. Sequences that are palindromic except for a few mismatches in base pairing are called near-palindromes. If the complementary portions are separated by a gap sequence, the sequence is referred to as inverse repeat 3. When a palindromic sequence is folded at its midpoint, the base pairs (bp) on the two halves are complementary. 1, the sequence on the positive strand 5′-GACA|TGTC-3′ is a palindrome since GACA is followed by its complement CTGT, but appearing in reverse order as TGTC 2. In DNA, palindromes are defined as a sequence of nucleotides that are followed by its complement sequence appearing in reverse order 1. The catalog of palindromes in the reference genome and 1000 Genomes is being made available here with details on their variations in each individual genome to serve as a resource for future and retrospective whole-genome studies identifying statistically significant palindrome variations associated with diseases or traits and their roles in disease mechanisms. The analysis of disease/trait-associated single-nucleotide polymorphisms in palindromic regions showed that disease-associated risk variants are 14 times more likely to be present in palindromic regions than in other regions. We found that ~30% of the palindromes exhibit variation, some of which are caused by rare variants. In this work, we computed and analyzed the palindromic sequences in the human genome and studied their conservation in personal genomes using 1000 Genomes data. Palindromic mutations are linked to many human diseases, such as neuronal disorders, mental retardation, and various cancers. Palindromes are distributed throughout the human genome and play significant roles in gene expression and regulation. A palindrome in DNA is like a palindrome in language, but when read backwards, it is a complement of the forward sequence effectively, the two halves of a sequence complement each other from its midpoint like in a double strand of DNA.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
